Sproul Lab Research
Current Projects
Mechanisms of Rapid Genome Evolution

Large fractions of the genome in most organisms are made up of repetitive DNA elements. Repetitive elements encompass the fastest evolving compartments of genomes and account for major variation in genome size, content, and organization, even among closely related species. Rapid repeat turnover can have major consequences for genome structure, regulation, and phenotypic change, however the mechanisms underlying rapid turnover remain poorly understood. We have characterized instances of rapid genome evolution in a group of nine closely related Bembidion ground beetle species. In this group, although gene-containing regions of genomes show modest variation across species, repetitive genomic regions containing tandem repeats (e.g., satellite DNA) can account for wholesale turnover of >40% their genomes overall. The image above shows fluorescence in-situ hybridization experiments that reveal a major shift in the abundance and genomic locations of 28S ribosomal DNA-derived repeats between two closely related ground beetle species. In the lower species, 28S rDNA is the source sequence of rDNA-like satellite DNA sequences which have subsequently proliferated throughout the genome. Variation in this single repetitive sequence accounts for >17% divergence in overall genome content between the two species. We are investigating the molecular mechanisms underlying rapid genome evolution in this group using a combination of long-read genome sequencing and assembly, RNA sequencing, and fluorescence in-situ hybridization experiments. We eventually plan to test whether rapid genome evolution is linked to reproductive barriers among species.
Repetitive Element Dynamics in Insect Genomes

Repetitive elements have been understudied for decades due to technical and computational challenges associated with their sequencing and assembly. They are particularly understudied in diverse groups of organisms that don’t have a history of genomics research. In addition, we know little about how repeat dynamics shape genome evolution in naturally evolving species across changing ecological landscapes. We are working to extend repetitive DNA genomics to the biodiversity genomics community investigating repetitive element dynamics across phylogenetic scales in insects. The above figure (generated as part of a collaborative paper here) reports repetitive element dynamics from 600 insect species and reveals many compelling avenues for future research. Studying repeats in a comparative evolutionary framework that spans populations, species, and diverse clades is a powerful lens for understanding patterns, processes, and mechanisms that underlie genome evolution, repeat biology, and the diversification of life.
Species delimitation, taxonomy, phylogeography

A thorough understanding of organisms, their traits, life histories, ecological interactions, species boundaries, and environments serves as a foundation for key observations that drive deep scientific discovery. We are passionate about organismal biology, taxonomy, and natural history collections. We have ongoing species delimitation and phylogeography projects in high elevation Bembidion ground beetles as well as stoneflies. We delimit species by integrating evidence from morphological, geographic, and genome-scale molecular data sets, including with novel approaches that use signal from repetitive DNA to help resolve species boundaries. This research takes us to beautiful alpine and sub-alpine habitats across western North America. Natural history collections are an indispensable source of specimens and locality data that supplement new collecting efforts. In addition to advancing discovery and documentation of biodiversity in focal taxonomic groups, the knowledge of population structure and species boundaries gained from this work informs projects on evolutionary processes and molecular mechanisms of rapid genome evolution described above.
Bioinformatic tool development
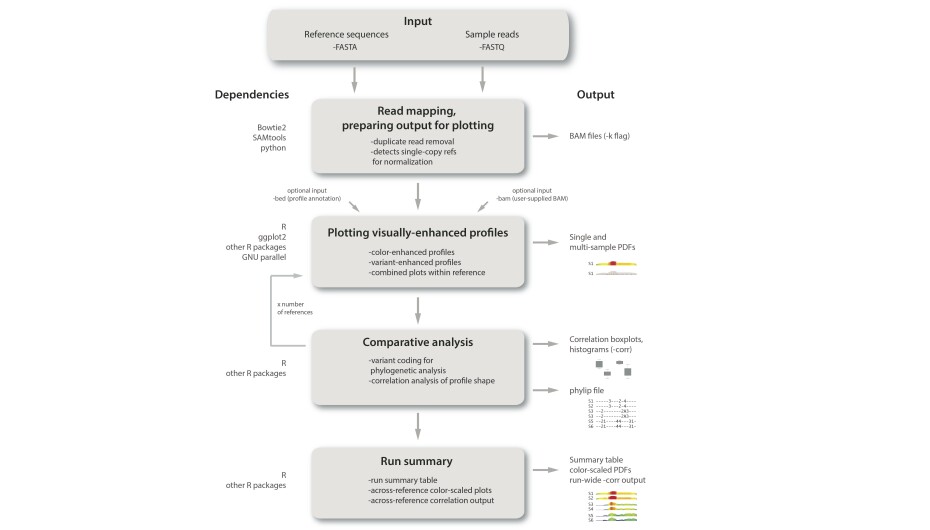
The technologies for generating molecular data sets continue to improve and diversify, as do software tools for analyzing the resulting data. In this dynamic data landscape, there are many opportunities for software tool development, especially in relatively young areas of research such as biodiversity-scale repeat genomics. In a past project led by University of Rochester undergrads Sherif Negm and Anya Greenberg, we developed RepeatProfiler, a pipeline for visualization and comparative analysis of repetitive DNA profiles. We also have a new pipeline nearing completion called SatHomologs that establishes homology of satellite DNAs within and across species. We routinely write less formal scripts and computational pipelines to automate analysis workflows on supercomputing clusters. The lab has coding expertise in Python, R, and Bash. With each project we are on the lookout for tool development opportunities that will help us ask new questions and accelerate high throughput repeat analysis in diverse clades.